As you use 23andMe's tools, you will find a few basic concepts coming up again and again. Our Genetics Guides are a useful starting place for people who are new to the science behind our tools, and a valuable reference for those who haven't encountered it recently.
- What is DNA?
- What is a chromosome?
- What is a SNP?
- What is a gene?
- What is recombination?
- What is mitochondrial DNA (mtDNA)?
- What is the Y chromosome (a.k.a. Why the Y)?
- What is special about the X chromosome?
- What is the difference between genetic distance and physical distance?
What is DNA?
DNA stands for DeoxyriboNucleic Acid, and it is a type of molecule found in most of the cells in your body. DNA is special because it encodes the basic instructions for making all the different types of cells in your body. In addition to containing the blueprint for muscle, brain, or blood cells, DNA has the instructions for how those cells work together.
How does a molecule encode anything? One molecule of DNA is actually a long string of many individual building blocks called nucleotides. These nucleotides come in different types depending on the base attached to them. There are four different bases: adenine (A), thymine (T), guanine (G), and cytosine (C). Using this simple “alphabet”, DNA can encode large amounts of information. Machinery in our cells can read the bases and decode this information to make all the pieces necessary for building and maintaining the cells of the body.
In the cell, DNA is generally stored in its most stable form: two strands of DNA wrapped around each other to form the famed double helix.

Each of these strands of DNA is composed of a series of nucleotides, and each base matches up with a preferred partner — these pairs are called ‘base pairs.’ An A on one strand always pairs with a T on the other, and likewise, G always pairs with C. Because of this pairing, we say that the two strands in a DNA molecule are “complementary”.
Humans share nearly identical DNA. But, with ~3 billion bases in the human genome, there is a substantial amount of variation within our species. Each base can be represented by a single letter (A, T, C, or G), and there are ~10 million positions in the genome where large numbers of people have different letters from one another.
What is a chromosome?
The human genome, which contains all the genetic material needed to make a person, is over three billion base pairs in length. If you stretched out the DNA from just one human cell, it would be almost six feet long! All that DNA isn’t in one long strand, though. Anyone who’s untangled a jump rope knows that a string that isn’t neatly coiled up gets pretty messy.
Our DNA is stored in 46 pieces called chromosomes. Each chromosome consists of DNA and the proteins that keep it organized into tightly wound coils. There are two types of chromosomes: sex chromosomes, and autosomes. The X and Y chromosomes are the sex chromosomes, and they determine whether you’re biologically female or male. Women have two X chromosomes, while men have an X and a Y (although there are exceptions to this rule). All the other chromosomes, which both men and women have, are called autosomes.
Our 46 chromosomes are divided up into pairs, so every chromosome has a partner, or homolog. Homologous chromosomes are very similar to each other and carry the same sets of genes, though those genes may vary a little between chromosome pairs. They’re like two copies of the same recipe with slightly different ingredients. For every homologous pair, one chromosome comes from your mother and the other from your father, which is why you’re a blend of both your parents.

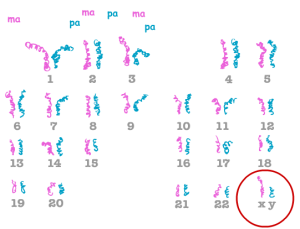
The above image shows the chromosomes for a normal human male: 22 pairs of autosomes, and one pair of sex chromosomes (one X and one Y). We have 46 chromosomes total, or 23 pairs of homologous chromosomes.
What is a SNP?
Humans share nearly identical DNA. But, with ~3 billion bases in the human genome, there is a substantial amount of variation within our species. Each base can be represented by a single letter (A, T, C, or G), and there are ~10 million positions in the genome where large numbers of people have different letters from one another. The locations with differences, called SNPs, are just one way each of us is genetically unique.
SNP (pronounced "snip") stands for Single Nucleotide Polymorphism. While the full name may be a mouthful, all it really describes is a single base position in our DNA that is variable in the general population.
For example, let’s say in 90% of the population, the DNA sequence along a certain stretch of chromosome 2 is ATGCCCGT and in 10% of the population, the sequence along that same stretch is ATGCACGT. The sequences are identical except at the fifth base, which is C in 90% of the population, and A in the other 10%. Because there are two different versions, C and A, at a single site, we call this location a single nucleotide polymorphism.
Some SNPs influence the function of a gene—how it is turned on or off, where it is read and used, or how well it does its job. If a gene plays an important role in the body, the two versions of a SNP may have noticeably different physical effects: changes in one’s phenotype. The effects of these causative (or functional) SNPs may be beneficial, as in the case of lactase persistence (the ability to digest milk as an adult); neutral, as in the case of hair color; or harmful, as in Tay-Sachs disease. The benefit or harm of one version or the other can even change depending on one’s environmental surroundings.
Other SNPs appear to make no functional difference at all, or at least none that scientists have identified yet. Regardless of whether a SNP has an obvious impact, it can be useful as a genetic marker. If a SNP is located very close to an unidentified genetic change that does have a physical effect, that SNP is linked to the effect, even if it doesn’t directly cause it. The SNP is then useful as a “landmark”, similar to how people describe locations. You may not know where the local hardware store is, but if you know it’s within a block of the pharmacy you went to the other day, you will likely be able to find it.
It is estimated that there are around 10 million SNPs in the human genome. That’s 100 times more than the hairs you have on your head! Because there are so many variable positions in the human genome, scientists often do not know which genetic change is responsible for which genetically-determined characteristic or disease. Many of today’s common diseases, such as obesity and heart disease, are quite complex, with multiple genetic and environmental components.
How do SNPs come to be? Every time your body makes a new cell, all your DNA has to be carefully copied so that the new cell has its own full set of chromosomes. Once in a very long while, the DNA copying machinery makes a mistake, and a single base may get changed. We call this change a mutation. If this mutation gets passed on to children and persists in the population, that location becomes a SNP.
What is a gene?
A gene is a segment of DNA that encodes a function. For example, there are genes that contain the instructions for making amylase (an enzyme in your saliva that helps break down starchy food), collagen (which forms much of the connective tissue in our bodies), and hemoglobin (the substance in our red blood cells that carries oxygen around).
Our cells have special machinery to read and translate DNA into products, often proteins. In the "genetic code," three consecutive DNA bases code for one amino acid, which is the basic building block for proteins. Proteins, in turn, give physical structure to our bodies, perform metabolic functions, etc.
For example, the DNA sequence AAA tells the cell machinery to add the amino acid lysine to a growing protein chain, whereas CGC codes for the amino acid arginine. In general, one gene encodes one protein. Changes to a gene's code can cause changes in the protein the gene codes for.
The human genome contains roughly 20,000 genes. It is the individual differences in these 20,000 genes, and the corresponding differences in their protein products, that make each of us unique.
Our genes are carried on long stretches of DNA called chromosomes, and each chromosome contains hundreds or thousands of genes. Because we have two copies of each chromosome (one from each parent), we have two copies of every gene.
What is recombination?
Reproduction is important for the evolutionary health of any species. For most of the chromosomes, children inherit one copy of each gene from their mothers and one copy from their fathers. Mixing and matching different versions of genes from our ancestors contributes to genetic diversity, which may enable us to adapt to unexpectedly stressful environments.
Recombination occurs during the production of sperm or eggs cells. This process allows the chromosomes you inherit from your mother to carry segments from both of her parents. The same is true for the chromosomes you inherit from your father and for the chromosomes they inherited from their parents—and so on back up your genealogical tree. If it weren't for recombination, each chromosome you inherited from your mother would have come entirely from either your grandmother or from your grandfather.
During the production of sperm or eggs, the homologous pairs of chromosomes cross at certain points and switch genetic material. As a result, the newly formed chromosomes are mixtures of the two originals.

If this is difficult to understand, try this. Imagine two hands of cards, one with only spades, the other with only hearts. Each hand represents one chromosome from the homologous pair. In the figure below, each hand has the cards Ace through 5, which represent genes on the chromosomes.

If you were to exchange the Aces and the 3s and 4s, you would have two new hands, each with the Ace through 5 but with a mixed set of suits. This exchange is similar to that of genetic recombination.

Recombination only occurs between homologous chromosomes, or chromosomes that come in pairs. However, there are two regions of the genome that only exist as one copy: the mitochondrial genome and part of the Y chromosome.
Since the Y chromosome doesn’t match up with the X chromosome completely, only the tips of the Y and X chromosomes recombine with one another. The tips of the Y chromosome that could recombine with the X chromosome are referred to as the pseudoautosomal regions. The rest of the Y chromosome is passed on to the next generation intact.
What is mitochondrial DNA (mtDNA)?
Small structures called mitochondria reside in almost every cell in your body. Within each of these structures is a tiny circular genome. We call the DNA of this genome "mitochondrial DNA" or "mtDNA" for short.
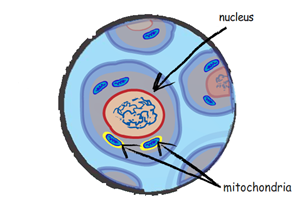
Unlike the rest of your genome, mtDNA is only passed on from mother to child; mtDNA inheritance is "maternal," tracking your ancestry through your mother, your mother's mother, your mother's mother's mother, and so on.
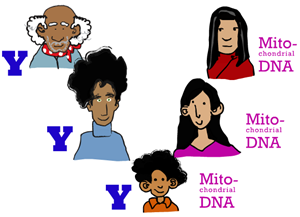
What is the Y chromosome (a.k.a. Why the Y)?
The Y chromosome is a sex chromosome found in males. It is passed from father to son. Compared to the other chromosomes, the Y chromosome does not contain many genes - it has almost 30 times fewer genes than the X chromosome. Most of the genes on the Y have male-specific functions, such as male sex determination or male fertility.
Unlike the other chromosomes, there is generally only one copy of the Y per cell. But it isn't left without a dance partner: during recombination the Y chromosome pairs up with the X. While other chromosomes recombine along their entire length, only the tips of the Y and X chromosomes recombine. The tips of the Y chromosome that could recombine with the X chromosome are referred to as the pseudoautosomal region. The rest of the Y chromosome is passed on to the next generation intact.
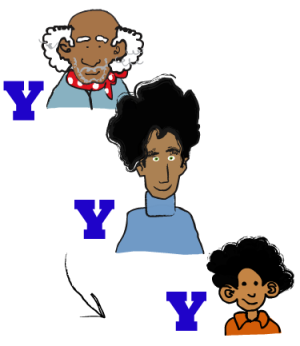

Because most of a son's Y chromosome is inherited only from his father, the Y chromosome is very useful for tracing geographic ancestry. The Y chromosomes of all living males are related through a single male ancestor who lived over 100,000 years ago. By looking at the geographic distribution of sets of closely related Y chromosome lineages (called haplogroups), we learn how our ancestors migrated throughout the world.
The Y differs from other regions of the genome because only males pass on Y chromosomes each generation (and not even all males, since not all men have sons). For this reason the Y chromosome is particularly susceptible to what is known as genetic drift, a process in which the frequencies of genetic variants can vary over time. These changes are more likely to occur because of population-level events such as migrations, new settlements, and replacement. And since the Y chromosome does not recombine, the history of the lineages affected by these events is preserved through the generations. As a result, Y chromosome variation has a strong geographic pattern.
What’s the difference between genetic distance and physical distance?
Physical distance along a chromosome is measured in terms of the number of base pairs (bp) or megabase pairs (Mb), where 1 Mb = 1000 bp. Genetic distance reflects recombination, and is measured in terms of centimorgans (cM). One cM is equivalent to the number of bp over which a single crossing over event is expected every 100 generations (so a 1% chance per generation). Genetic distance is irregular across chromosomes as well as along a chromosome. The measurement in centimorgans takes into consideration both the physical distance and the number of crossing-over events that typically occur in that region. This irregularity means that you cannot simply add and subtract centimorgans without knowing details about how the typical number of crossing over events changes along that particular segment.
What is special about the X chromosome?
Men and women inherit the X chromosome differently. Men inherit their X chromosome from their mother, while women inherit an X chromosome from each parent. Since men inherit this chromosome differently than women, only certain ancestors could have contributed to the segments of DNA located on your X chromosome.
This means that if you are sharing a segment of DNA with someone on the X chromosome, you can immediately dismiss certain ancestors as your recent common ancestor.
If you are a male matching someone on the X chromosome, you can immediately dismiss any ancestors on your father's side of the family. Your common ancestor can only be located on your mother's side of the family. Below you can see your recent ancestors (highlighted in blue) that could have contributed to your X chromosome.

If you are female matching someone on the X chromosome, you can dismiss the ancestors located on father/son lines. As an example, you can dismiss your paternal grandfather's family since your father inherited his X chromosome from his mother. Below you can see your recent ancestors (highlighted in yellow) that could have contributed to your X chromosome.
